论文分析-百度eHealth:Building Chinese Biomedical Language Models via Multi-Level
用Xmind画的思维导图,导出的markdown缺少图片,可以看文末的图片更为完整
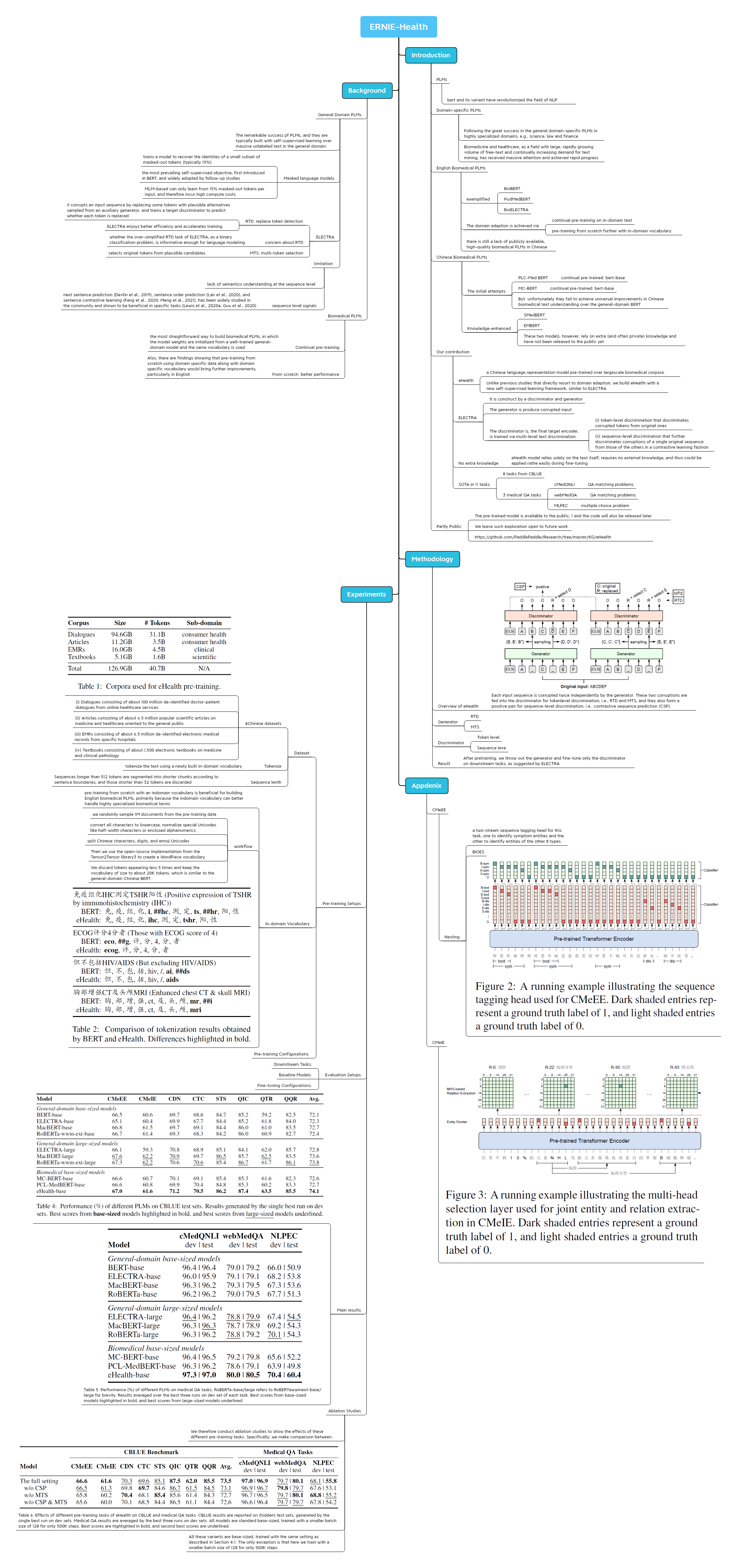
ERNIE-Health
https://arxiv.org/abs/2110.07244
Introduction
PLMs
- bert and its variant have revolutionized the field of NLP
Domain-specific PLMs
- Following the great success in the general domain-specific PLMs in highly specialized domains, e.g., science, law and finance
- Biomedicine and healthcare, as a field with large, rapidly growing volume of free-text and continually increasing demand for text mining, has received massive attention and achieved rapid progress
English Biomedical PLMs
exemplified
- BioBERT
- PudMedBERT
- BioELECTRA
The domain adaption is achieved via
- continual pre-training on in-domain text
- pre-training from scratch further with in-domain vocabulary
there is still a lack of publicly available, high-quality biomedical PLMs in Chinese
Chinese Biomedical PLMs
The initial attempts
PLC-Med BERT
- continual pre-trained: bert-base
MC-BERT
- continual pre-trained: bert-base
But unfortunately they fail to achieve universal improvements in Chinese biomedical text understanding over the general-domain BERT
Knowledge-enhanced
- SMedBERT
- EMBERT
- These two models, however, rely on extra (and often private) knowledge and have not been released to the public yet
Our contribution
eHealth
- a Chinese language representation model pre-trained over largescale biomedical corpora.
- Unlike previous studies that directly resort to domain adaption, we build eHealth with a new self-supervised learning framework, similar to ELECTRA
ELECTRA
- It is construct by a discriminator and generator
- The generator is produce corrupted input
The discriminator is, the final target encoder, is trained via multi-level text discrimination.
- (i) token-level discrimination that discriminates corrupted tokens from original ones
- (ii) sequence-level discrimination that further discriminates corruptions of a single original sequence from those of the others in a contrastive learning fashion
No extra knowledge
- eHealth model relies solely on the text itself, requires no external knowledge, and thus could be applied rathe easily during fine-tuning
SOTA in 11 tasks
- 8 tasks from CBLUE
3 medical QA tasks
cMedQNLI
- QA matching problems
webMedQA
- QA matching problems
MLPEC
- multiple choice problem
Partly Public
- The pre-trained model is available to the public, 1 and the code will also be released later.
- We leave such exploration open to future work.
- https://github.com/PaddlePaddle/Research/tree/master/KG/eHealth
Background
General Domain PLMs
- The remarkable success pf PLMs, and they are typically built with self-supervised learning over massive unlabeled text in the general domain.
Masked language models
- trains a model to recover the identities of a small subset of masked-out tokens (typically 15%)
- the most prevailing self-supervised objective, first introduced in BERT, and widely adopted by follow-up studies
- MLM-based can only learn from 15% masked-out tokens per input, and therefore incur high compute costs
ELECTRA
RTD: replace token detection
- it corrupts an input sequence by replacing some tokens with plausible alternatives sampled from an auxiliary generator, and trains a target discriminator to predict whether each token is replaced
- ELECTRA enjoys better efficiency and accelerates training.
concern about RTD
- whether the over-simplified RTD task of ELECTRA, as a binary classification problem, is informative enough for language modeling
MTS: multi-token selection
- selects original tokens from plausible candidates.
limitation
- lack of semantics understanding at the sequence level
sequence level signals
- next sentence prediction (Devlin et al., 2019), sentence order prediction (Lan et al., 2020), and sentence contrastive learning (Fang et al., 2020; Meng et al., 2021), has been widely studied in the community and shown to be beneficial in specific tasks (Lewis et al., 2020a; Guu et al., 2020).
Biomedical PLMs
Continual pre-training
- the most straightforward way to build biomedical PLMs, in which the model weights are initialized from a well-trained general-domain model and the same vocabulary is used
From scratch: better performance
- Also, there are findings showing that pre-training from scratch using domain specific data along with domain specific vocabulary would bring further improvements, particularly in English
Methodology
Overview of eHealth
- Each input sequence is corrupted twice independently by the generator. These two corruptions are fed into the discriminator for tokenlevel discrimination, i.e., RTD and MTS, and they also form a positive pair for sequence-level discrimination, i.e., contrastive sequence prediction (CSP).
Generator
- RTD
- MTS
Discriminator
- Token level
- Sequence leve
Result
- After pretraining, we throw out the generator and fine-tune only the discriminator on downstream tasks, as suggested by ELECTRA
Experiments
Pre-training Setups
Dataset
4Chinese datasets
- (i) Dialogues consisting of about 100 million de-identified doctor-patient dialogues from online healthcare services
- (ii) Articles consisting of about 6.5 million popular scientific articles on medicine and healthcare oriented to the general public
- (iii) EMRs consisting of about 6.5 million de-identified electronic medical records from specific hospitals
- (iv) Textbooks consisting of about 1,500 electronic textbooks on medicine and clinical pathology.
Tokenize
- tokenize the text using a newly built in-domain vocabulary
Sequence lenth
- Sequences longer than 512 tokens are segmented into shorter chunks according to sentence boundaries, and those shorter than 32 tokens are discarded
In-domain Vocabulary
- pre-training from scratch with an indomain vocabulary is beneficial for building English biomedical PLMs, primarily because the indomain vocabulary can better handle highly specialized biomedical terms
workflow
- we randomly sample 1M documents from the pre-training data
- convert all characters to lowercase, normalize special Unicodes like half-width characters or enclosed alphanumerics
- split Chinese characters, digits, and emoji Unicodes
- Then we use the open-source implementation from the Tensor2Tensor library3 to create a WordPiece vocabulary
- We discard tokens appearing less 5 times and keep the vocabulary of size to about 20K tokens, which is similar to the general-domain Chinese BERT.
Pre-training Configurations.
Evaluation Setups
- Downstream Tasks.
- Baseline Models.
- Fine-tuning Configurations.
Main results
- Table 5: Performance (%) of different PLMs on medical QA tasks. RoBERTa-base/large refers to RoBERTawwmext-base/large for brevity. Results averaged over the best three runs on dev set of each task. Best scores from base-sized models highlighted in bold, and best scores from large-sized models underlined.
Ablation Studies
- We therefore conduct ablation studies to show the effects of these different pre-training tasks. Specifically, we make comparison between:
- Table 6: Effects of different pre-training tasks of eHealth on CBLUE and medical QA tasks. CBLUE results are reported on (hidden) test sets, generated by the single best run on dev sets. Medical QA results are averaged by the best three runs on dev sets. All models are standard base-sized, trained with a smaller batch size of 128 for only 500K steps. Best scores are highlighted in bold, and second best scores are underlined.
- All these variants are base-sized, trained with the same setting as described in Section 4.1. The only exception is that here we train with a smaller batch size of 128 for only 500K steps.
Appdenix
CMeEE
Nesting
- a two-stream sequence tagging head for this task, one to identify symptom entities and the other to identify entities of the other 8 types.
- BIOES
CMeIE
Mind Map